
The goal of the research in my laboratory is to understand fundamental paradigms that govern metabolic network design. We use a bacterial model system and emphasize a biochemical genetic approach that utilizes in vivo analyses to inform the design of in vitro experiments. This goal has led us into probing the role of non-enzymatic chemistry and metabolic crosstalk in metabolism, in addition to engaging collaborators with metabolomics and mathematical modeling expertise. Currently the work in the lab is in two general areas: 1) Understanding the Rid system of endogenous metabolite stress, and 2) Exploring metabolic integration and redundancy using a combination of classical and global approaches.
Students from my laboratory have strong training in classical and molecular genetics, particularly as applied to metabolic questions. In addition, they utilize standard biochemical and molecular biological approaches. I encourage students to think logically about big biological questions and how to tease them apart. I strive to train students to think beyond linear pathways to appreciate the integrated nature of metabolism and the inherent chemistry that governs cellular processes.
I am interested in furthering our understanding of the structure and integration of bacterial metabolic networks, how they respond to perturbation, and how metabolism is integrated into specialized organismal capabilities. The overall goal of the research in my laboratory is to understand fundamental paradigms that govern metabolic network design. We use a bacterial model system and emphasize a biochemical genetic approach that utilizes in vivo analyses to inform the design of in vitro experiments. In total, we strive to think rigorously and creatively about metabolism in an effort to understand the true metabolic potential of any organism.
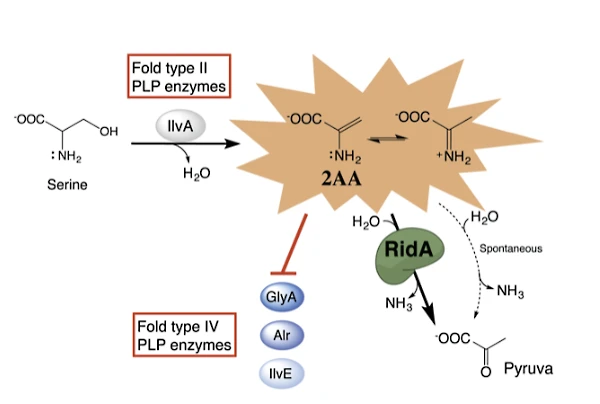
My laboratory continues to expand our understanding of the RidA protein that was identified and characterized as an enamine deaminase by my group. We showed that the enamine 2-aminoacrylate, which is an obligate intermediate in some PLP-dependent reactions, can damage cellular components if it accumulates. The RidA protein, a member of the Rid superfamily (previously YjgF/YER057c/UK114) is responsible for deaminating 2AA to prevent its accumulation, making it a novel stress response protein. Our findings have opened new directions of study in the lab. Immediate questions include; which enzymes generate enamine stressors? Which enzymes are targets of the damage? What are the biochemical consequences of lacking RidA in various organisms?
We have expanded our work to define the consequences of lacking RidA in Saccharomyces cerevisiae, Pseudomonas aeruginosa and Campylobacter jejuni. The differences in metabolic network structure between organisms has been emphasized by the different responses each has to the lack of this critical enzyme. New projects address the role of the seven non-RidA subfamilies (Rid1-7, RutC) of the Rid superfamily. Current data suggests these subfamilies, found only in prokaryotes, are components of diverse metabolic pathways that provide niche specific fitness.
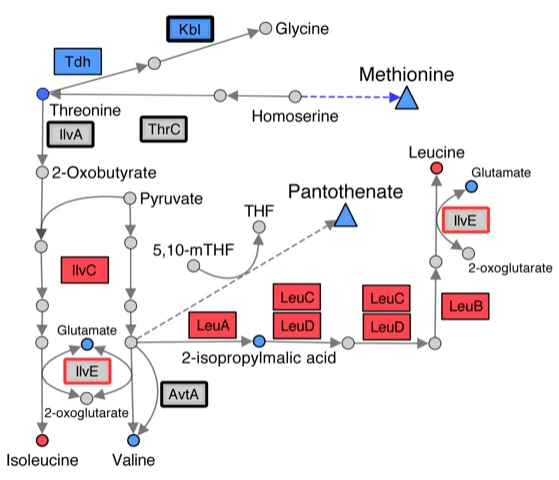
By virtue of the selective pressure exerted through millions of years of evolution, a living cell is likely to be the most well-tuned complex system in existence. Much of the research in the laboratory uses classical approaches and emerging technologies to better understand the molecular details of metabolic processes, and identify the mechanisms used to integrate these processes into a productive physiology. This approach continues to lead us to identifying the role(s) of previously unknown proteins, and contributes to our effort to define metabolic structure using global approaches. Recent addition of collaborators with metabolomics and mathematic modeling expertise has invigorated these studies, which have been the long-term interest of the lab. Current efforts utilize NMR metabolomics approaches in collaboration with the Edison laboratory at UGA. These metabolomics approaches can help accelerate the description and understanding of complex metabolic and physiological states of an organism. As such these approaches help complement the biochemical and genetic approaches that have identified the metabolic framework and raised broad global questions over the years. Moving forward we expect to incorporate metabolomic studies into our analysis of how the lack of RidA, or other central components, perturbs the metabolic network structure of various model organisms.
My laboratory consistently has a strong and vibrant group of students that enjoy thinking about science and developing problem solving skills. We emphasize the development of models and the design of experiments to test them. Graduate and undergraduate students form the critical component of the lab community, which over time has also included, high school students, post-doctoral fellows and technicians. The laboratory emphasizes “big picture” thinking and thrives on complex questions that require one to integrate multiple variables and delve into the literature to recall the basics of metabolism, into which new data must be incorporated. Students from my laboratory have strong training in classical and molecular genetics particularly as applied to metabolic questions. They also utilize standard biochemical and molecular biological approaches. The direction of the research ensures that students will become familiar with the global metabolomics and mathematic modeling approaches used in Systems Biology. I strive to train students to think beyond linear pathways to appreciate the integrated nature of metabolism and the inherent chemistry that governs cellular processes.
Tetrahydrofolate levels influence 2-aminoacrylate stress in Salmonella enterica.
Shen W, Downs DM.
Fulton RL, Downs DM.
Fulton RL, Downs DM.
Lab and Shipping Address
University of Georgia
Department of Microbiology
527 Biological Sciences Building
120 Cedar Street
Athens, GA 30602
Phone & Email
Office Ph: (706) 542-9573
Lab Ph: (706) 542-9579
Fax: (706) 542-2674
dmdowns@uga.edu